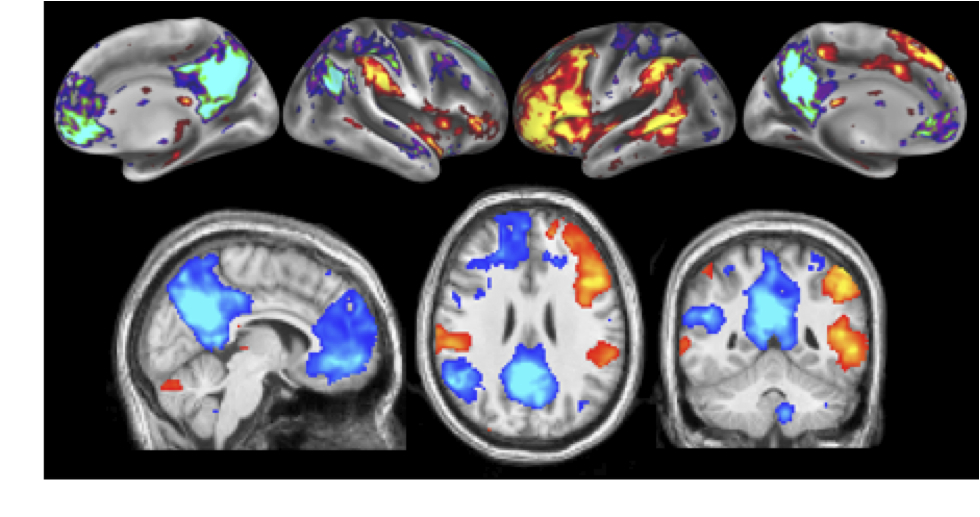
Resting-state functional magnetic resonance imaging (FMRI) has become a powerful tool for the study of functional networks in the brain. Even "at rest", the brain's different functional networks spontaneously fluctuate in their activity level; each network's spatial extent can therefore be mapped by finding temporal correlations between its different sub-regions. Current correlation-based approaches measure the average functional connectivity between regions, but this average is less meaningful for regions that are part of multiple networks; one ideally wants a network model that explicitly allows overlap, for example, allowing a region's activity pattern to reflect one network's activity some of the time, and another network's activity at other times. However, even those approaches that do allow overlap have often maximised mutual spatial independence, which may be suboptimal if distinct networks have significant overlap. In this work we identify functionally distinct networks by virtue of their temporal independence, taking advantage of the additional temporal richness available via improvements in FMRI sampling rate. We identify multiple "temporal functional modes", including several that subdivide the default-mode network (and the regions anti-correlated with it) into several functionally distinct, spatially overlapping, networks - each with its own pattern of correlations and anti-correlations. These functionally-distinct modes of spontaneous brain activity are, in general, quite different from resting-state networks previously reported, and may have greater biological interpretability.
See a movie of TFM dynamics - each timepoint is one TR=0.8s. Different colours are different TFMs; within a TFM, darker clusters are anticorrelated with brighter clusters.
Queries on TFMs: Steve Smith
Queries on the WashU-UMinn HCP: David Van Essen
Queries on the accelerated, short-TR image acquisition: Kamil Ugurbil
Data usage acknowledgements: if you use these datasets in your work, we request that you cite the above paper (Smith et al., PNAS, 2012), and also acknowledge that "data were made available by the WU-Minn Human Connectome Project (1U54MH091657), funded by the 16 NIH Institutes and Centers that Support the NIH Blueprint for Neuroscience Research."
Download NIFTI versions: 21 TFMs / 21 RSNs.
The RSN and TFM datasets are also available in the SumsDB database. This includes whole-brain volume slices viewable using Caret5 software (and also online using WebCaret); see the SumsDB README file. The surface-based visualization of RSN and TFM data make use of the CIFTI files that combines surface vertices and subcortical grey-matter voxels into a single file format. These datasets are not yet viewable in WebCaret (work in progress). They can be downloaded for visualization using the Connectome Workbench, which is also a work in progress. Investigators interested in accessing a beta-test version of Connectome Workbench should contact David Van Essen.
The full 4D NIFTI datasets from this study are now available from the HCP website.