

Next: Segmentation of Brain MR
Up: Segmentation Using the HMRF-EM
Previous: Initial Parameter Estimation
Various experiments have been carried out to test the performance
of the HMRF-EM framework. An example is shown in the following
figures. Figure 2(a) shows a simulated 3-class image
sampled from an MRF model using the Gibbs sampler. The intensities
for the three classes are 30, 125 and 220 respectively. Figure
2(b)-(e) show the same images with added Gaussian noise
with standard deviation of 28, 47, 66, and 95. Because image
contrast is what we are most interested in for examining qualities
of an image, a measurement of the noise is more meaningful with
image contrast being taken into account. Thus we define a measure,
the noise-to-contrast ratio (NCR) as the following:
Thus, the NCRs of the four test images are 0.3, 0.5, 0.7 and 1.0,
respectively. Figure 2(f)-(k) show their intensity
histograms. Except for the first, each histogram exhibits severe
overlap. The true parameters for the test images are listed in
Table 1.
Figure 2:
Test images for parameter estimation. (a) the original
image; (b)-(e) noisy images with NCR 0.3, 0.5, 0.7, and 1.0;
(f)-(k) histogram of (b)-(e).
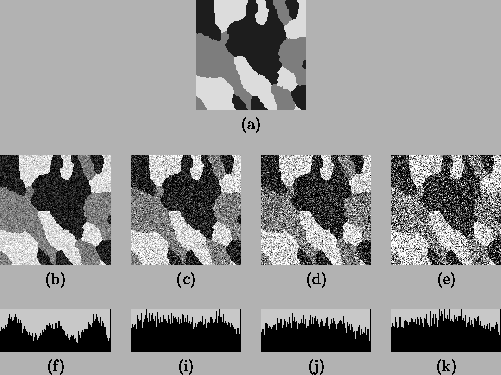 |
Table 1:
True model parameters of Figure
2(b)-(e).
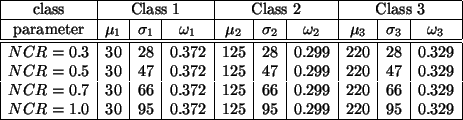 |
Table 2:
Initial parameter estimation using discriminant
measure-based thresholding.
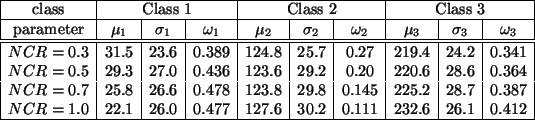 |
The discriminant measure-based thresholding method is then applied
to each of the four test images to estimate the initial
parameters. Table 2 shows the results. Comparing it
with Table 1, we can see that the estimates are
acceptable, especially when the noise level is low.
The standard FM-EM algorithm and the HMRF-EM algorithm are then
applied to the four test images until there is no significant
change in the value of the Q-function. To measure the
segmentation accuracy, we also define the misclassification ratio
(MCR), which is
The standard FM-EM algorithm only converges for the first image,
which has the lowest noise level (NCR=0.3). In this case, the
estimation results and the number of iterations K are shown in
Table 3. With those estimated parameters, we reconstruct
the histogram and obtain the segmentation, as shown in Figure
3. Note that, the parameter estimation is not accurate
when compared with their true values listed in Table 1.
Table 3:
Parameter estimation using the FM-EM algorithm
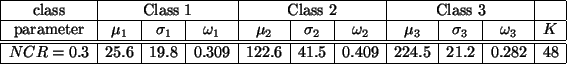 |
Figure 3:
Parameter estimation for Figure 2(b) using the
standard FM-EM algorithm. (a) the reconstructed histogram; (b) the
segmentation with MCR 5.82%.
 |
The HMRF-EM algorithm rapidly converges for all the four test
images. Table 4 and Figure 4 show the results.
Taking the true parameters shown in Table 1 as the
references and comparing the results from the two methods, it can
be seen that: (1) the HMRF-EM algorithm gives more accurate
estimates for parameters; (2) the HMRF-EM algorithm provides
automatic segmentation with much lower MCR.
Table 4:
Parameter estimation using the HMRF-EM algorithm
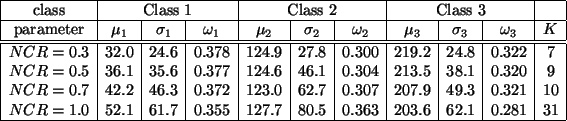 |
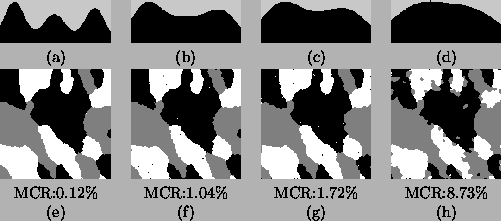
Figure 4:
Parameter estimation for Figure 2(b)-(e) using
the HMRF-EM algorithm. top row: the reconstructed histograms;
bottom row: the segmentations.
 |


Next: Segmentation of Brain MR
Up: Segmentation Using the HMRF-EM
Previous: Initial Parameter Estimation
Yongyue Zhang
2000-05-11


